Good evening everyone, I want to ask you a question, maybe you can help me. I am analyzing my data for the V3-V4 region (I have previously read Why do I have such a large distance matrix which tells me why these two regions should not be sequenced, however, these are the data that we have at the moment and therefore we are looking for some preliminary results). I have been doing the analyzes in an HPC cluster, however following the MiSeq tutorial in the OTUs section I have problems with the time it takes with at least 2 million sequences at least for what I think it should be in an HPC cluster. high performance. The command that I use is cluster.split but there comes a point where it hangs and does not advance any further, I have tried the other alternative of dist.seqs but this has taken much longer, for the execution I do it in a node of I work with 256GB of RAM and 256 processors, we also have access to reserve the use of GPUs in the cluster but apparently this does not speed up the process either.
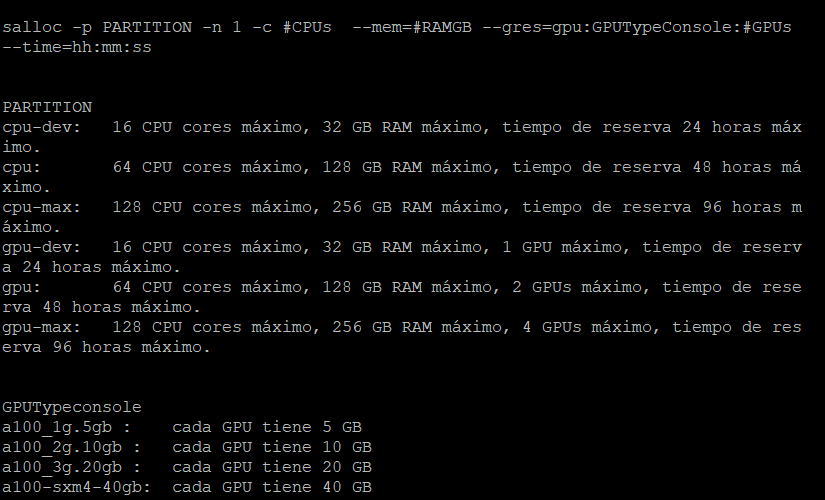
I would like to know if there is any way this can be set up so that the process of this stage progresses better. I leave you an image of the resources that we can reserve in the cluster